
A variety of layout algorithms are available, including cyclic, tree, force-directed, edge-weight, and yFiles Organic layouts. Node/edge/network), Desktop states (selected/hidden nodes and edges, window sizes), Properties, some plugin states, and Visual Styles. Cytoscape Session file includes networks, attributes (for It is called Cytoscape Session (.cys) file. All of settings, data files, and visualizations are packed in a session file. You can save your work by a single click. Networks, applying layouts, or image export. You can use your choice of programming languages to access Cytoscape core features, such as creating For example, if you have a network data generated byīioconductor, Cytoscape can load the file as a text table and you can export it in PSI-MI format for other bioinformatics tools or your own applications/scripts.įrom version 3.3, Cytoscape supports RESTful API for programmatic access. Since Cytoscape supports import/export standard file formats, you can easily put Cytoscape into your workflow. To develop new service clients for popular databases. IntAct, BioMart, and NCBI Entrez Gene are supported. This means Cytoscape can directly connect to the external public databases and imports network and annotation data. For example, input a set of custom annotation terms for your proteins, create a set of confidence values for your protein-protein interactions.Ĭytoscape works as a web service client. 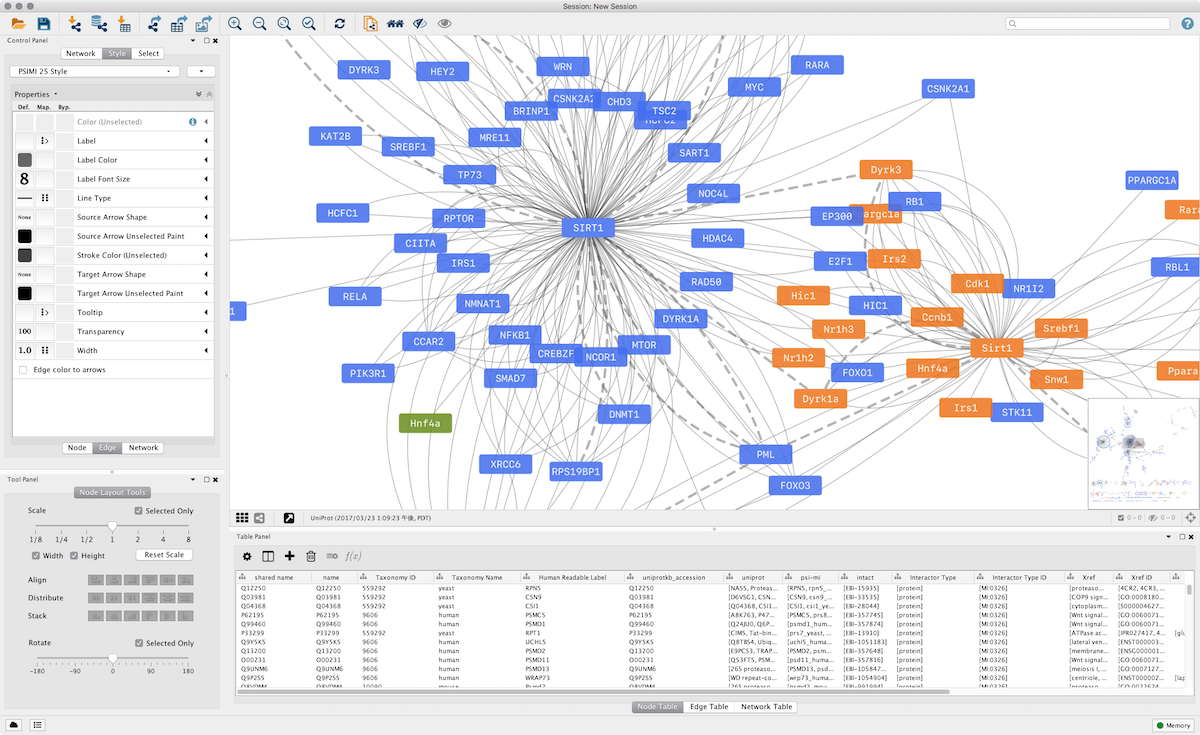
Using this feature, you can loadĪnd save arbitrary attributes on nodes, edges, and networks. 
Delimited text files and MS Excel™ WorkbookĪre also supported and you can import data files, such as expression profiles or GO annotations, generated by other applications or spreadsheet programs. Term, this library and other server-side applications replaces Java Web Start version of Cytoscape.Ĭytoscape supports a lot of standard network and annotation file formats including: SIF (Simple Interaction Format), GML , It is a pure JavaScript library and no plugin is required for the web browsers. Web Application Developers: You can use cytoscape.js as network data visualization engine for your web application. 3.xĪPI is stable enough to develop new Apps, but if you notice any design flaws or bugs, We recommend to use 3.x for new plugins development, as 2.x is desupported. Use v2.x if you have used v2.x in the past and rely on a plugin not supported as a v3.x app in the App Store – keep checkingĪpp Developers: Please port your 2.x Apps to 3.x series. Users, Casual Users, and Power Users: Use Cytoscape v3.x - v2.x has been desupported. Use this version unless you really need to use plugins only for 2.x series. New features will not be added by the core developers, and it may be hard to get answers to 2.x specific questions. This background is rendered by Cytoscape.js Ĭytoscape 2.x is a legacy version of Cytoscape. From Cytoscape 3.2, users can export network visualizations as interactive web application The Cytoscape core developer team is trying to improve interoperability of Cytoscape 3 and Cytoscape.js. It is designed to be a building block for complex data visualization Cytoscape 3 and cytoscape.js share design-level concepts, such as Visual Styles, but their code bases are completely independent to each other. Library for network visualization and analysis, and is NOT a complete webĪpplication. Please read roadmap page for further information.Ĭytoscape Web. Use the latest version of Cytoscape 3.x series. If you really need to use Plugins only available for 2.x, you may use Cytoscape 2.8.3, but otherwise, please As of May 2016, transition from Cytoscape 2.x to 3.x is complete, and 2.x is no longer supported. New features, including new user interfaces, advanced visualization functions, headless (command-line) distribution, RESTful API, and multiple rendering engine support, will be released for this It is a Java desktop application designed for large-scale It is designed for long-term maintainability and it replaced 2.x series. Cytoscape 3 is the mainstream version of Cytoscape with modular architecture. They may be developed by anyone using the Cytoscape open API based on Java™ technology and App community development is encouraged. Apps are available for network and molecular profiling analyses, new layouts, additional file format support, scripting, and connection with databases. Cytoscape core distribution provides a basic set of features for data integration, analysis, and visualization. Although Cytoscape was originally designed for biological research, now it is a general platform for complex networkĪnalysis and visualization. integrating these networks with annotations, gene expression profiles and other state data. Visualizing molecular interaction networks and biological pathways and 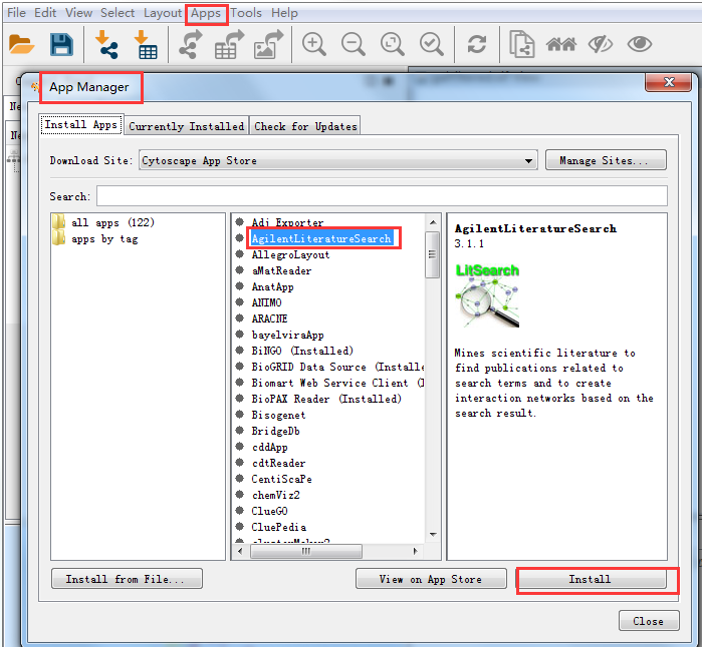
Cytoscape is an open source software platform for
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
